klone Users Storage Optimizations
Nam Pho
Director for Research Computingnote
There are steps you, as a researcher using klone, can do to limit the impact of whatever else is happening on the cluster on your individual workflows.
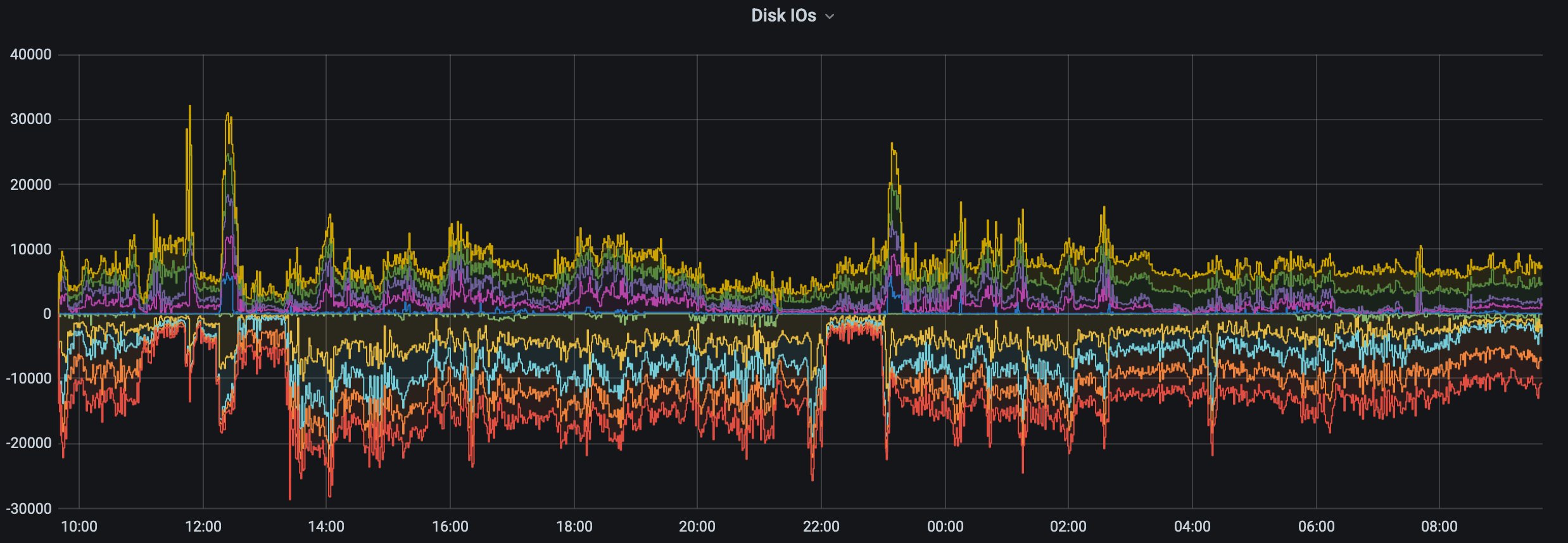
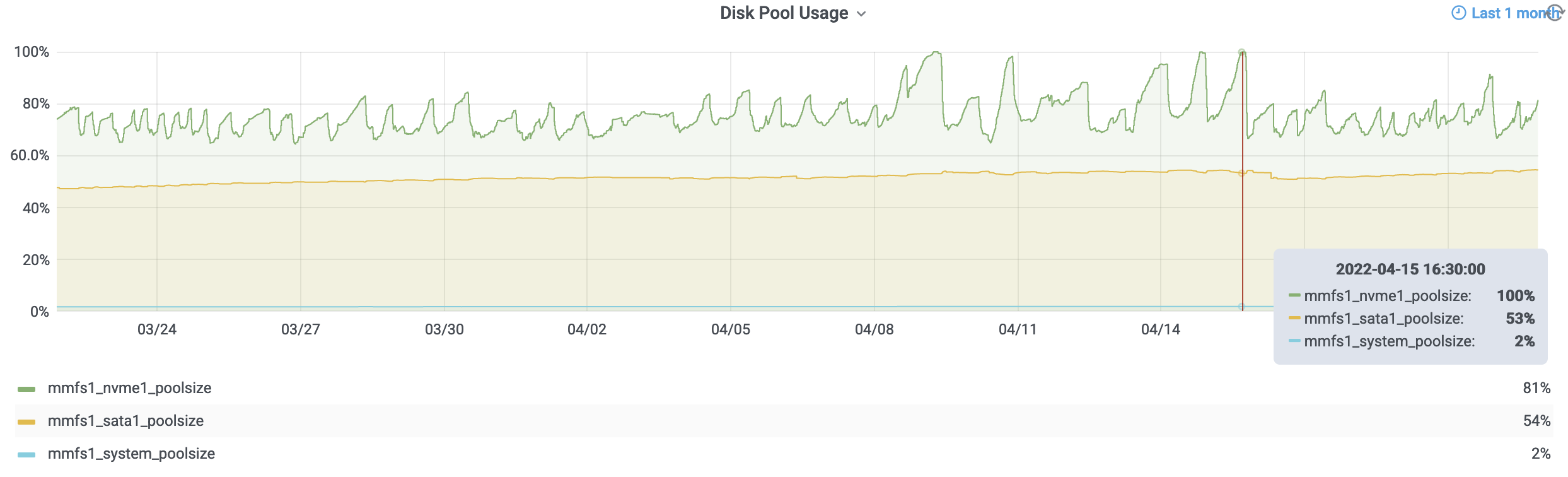
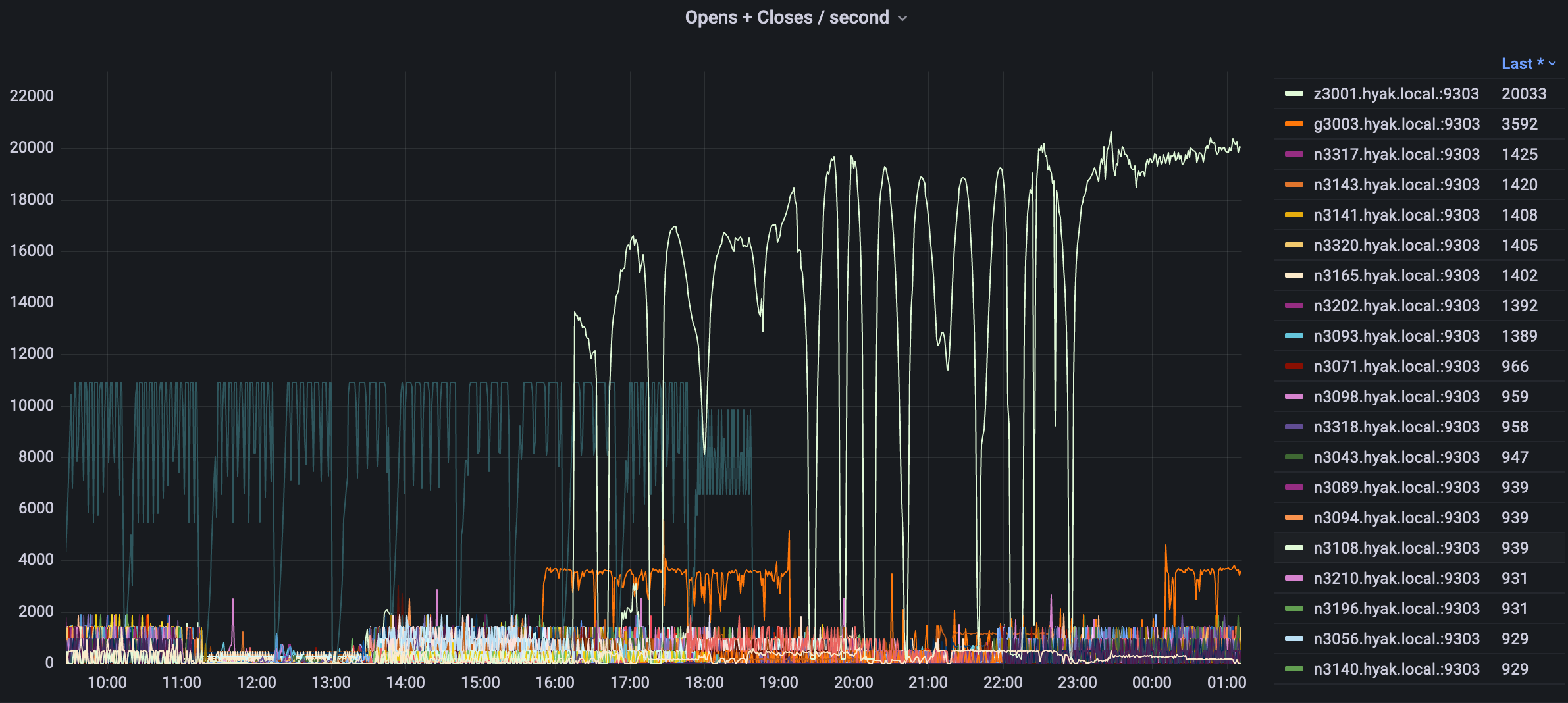
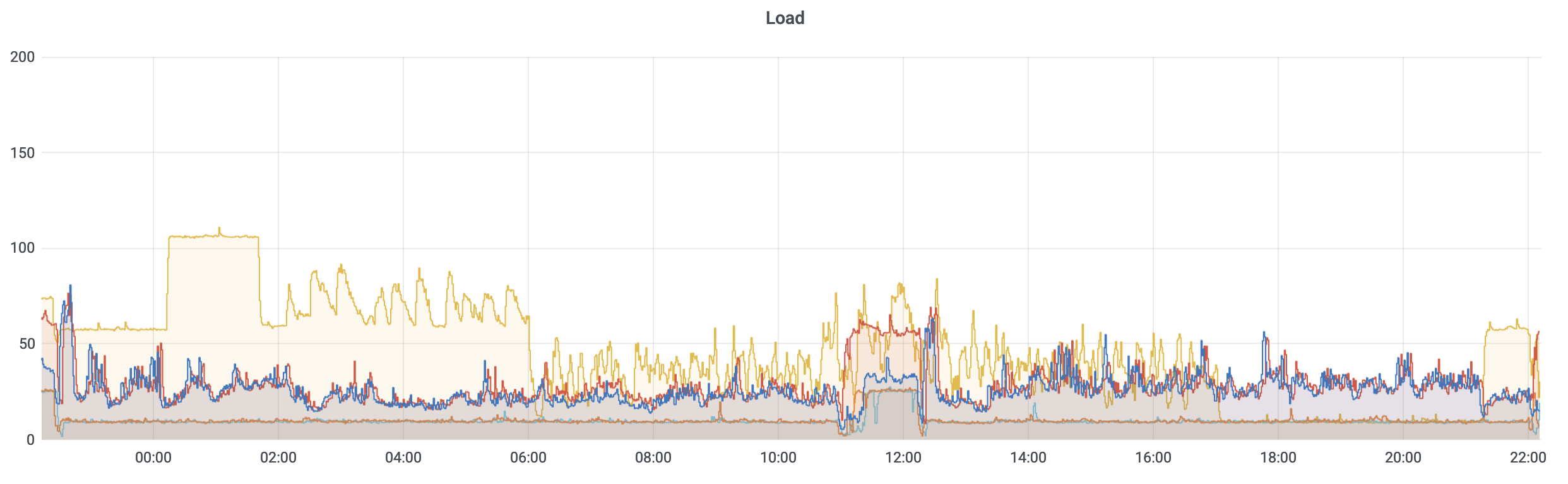
While some of what precipitated this conversation is the current state of the storage (i.e., mmfs1 or gscratch), there are several things you can do as a researcher to both reduce the load on gscratch as well as help insulate your jobs from cluster-wide storage slowdowns.
1. Use local node SSDs.#
Each node on the cluster has a local SSD drive with 350+ GB of space available for use by user jobs. This space is available only to jobs running on that node and all contents are purged when the users’ last job running on the node completes. It is mounted as /scr and /tmp (both paths go to the same place) on all the compute nodes.
If input data, Apptainer (Singularity) images, or other files used by your job will fit, copying those files to the SSD (via cp, rsync, etc.) once at the beginning of your job and reading them from there during the remainder of the job run results in less load on the central storage, helps insulate your job from any instances of central storage slowness, and can often result in better overall job performance.
Slurm has a command called sbcast [www] that is useful for efficiently copying files to all nodes used in a multi-node job as part of an sbatch script.
For files being written that need to be kept after the job run, it is generally best to write these directly to the central storage. Because new files are written directly to the very fast NVMe layer, such writes are less likely to impact overall storage performance. That said, it is still beneficial to write intermediate job files to the local SSD whenever possible.
2. Code for efficient file IO.#
While this can be a very complicated topic, a great deal of overall job performance can be gained by thoughtful and judicious use of file input-output (IO). Some general tips:
- Keep in mind that file access is orders of magnitude slower than memory access, and processes often have to completely "stop and wait" for disk IO operations to complete. Minimizing file IO operations, especially inside "inner loops" of programs can greatly speed up job completion, and helps to reduce load on the cluster central storage.
- Fewer, larger file IO operations are generally more efficient than multiple smaller file operations accessing the same data.
- When possible, store data in an efficient format such as HDF5 instead of many small files.
- "Open/read once, access many times" if job memory permits.
3. Containerize your environment.#
As mentioned above, minimizing the number of files you need to access can help reduce the number of input / output operations per second (IOPS) happening on the cluster. For example, a Python miniconda environment can create hundreds or even thousands of small files when you install different library dependencies. While Python is a common compute environment, this can be generalized to most other programs you may need. When you containerize your environment, this gets reduced to a single file. A brief introduction to Singularity (now called Apptainer) can be found here. As a side benefit, containerizing your environment–making it a single file–makes it much easier to move it around (see #1 above).
4. Stay under quota.#
Constantly hitting your inode (e.g., file) or block (e.g., number of GBs or TBs) quotas can cause extra storage slowness. If you need a bump on either please reach out to discuss your options. As a reminder you can us the hyakstorage command on klone to display current quota usage for all of your filesets as well as your home directory. Please note that this output is updated once an hour so it will take time to reflect any overages.
5. Report issues.#
While the Hyak team has an extensive monitoring and alerting framework in place to help us to proactively determine when things may be going wrong, not all causes of slow user experience are currently correlated to metrics. Furthermore, our team generally interfaces with the cluster in different ways than our users, so we may not be as equally exposed to any pains until it is reported to us. If you’ve run into a performance issue, please submit a ticket by emailing help@uw.edu. Please provide any symptoms you are observing, along with the date, timeframe, job IDs (if applicable), commands you are running with their full output, etc. If you don’t need or want a reply from us it is still helpful for us to hear from you, feel free to say "no response needed" or something along these lines so we know how to respond.
See also: